Chopin’s prelude No. 4 in E minor is widely venerated for its authentic musical depiction of despair. The haunting melody from gripping start to releasing finish has evoked profound emotions in listeners and players alike for over a century and a half. Examining the sheet music would illustrate how the piece is a constitution of hundreds of notes whose coordinated and delicate execution engenders its emotional valence. The piece is thus not merely the sum of its notes.
Our brain is also like a piano, where the music is our perceptions, memories, and emotions and the keys are the billions of neurons whose activity gives rise to such phenomena. This metaphor highlights a principal objective in neuroscience: if we ever hope to understand how the brain generates aspects of conscious experience, then we must read its sheet music. What is the orchestral firing of ensembles of neurons? Advances in neuroimaging have granted us access into this symphony, where we can literally see single-neuron activity as flickering balls of light. The field of optogenetics has paved the way to functionally interrogate these same neural circuits with light. What if we could read the sheet music with neuroimaging and then play the keys back to the brain? Could we manufacture emotion or sensation? This read-write capability requires fine spatiotemporal activation of identified neurons. This is what holographic optogenetics aims to accomplish.
In this article, I will share the fundamentals of neuroimaging and optogenetics and describe the optical technology being developed to activate specific populations of neurons. Finally, I will discuss how it is being used to write the music of the brain and play it back to itself.
Table of contents:
1. Fundamentals of Calcium Imaging: Reading the brain's sheet music
2. Fundamentals of Optogenetics: The molecular tools to write into neural circuits
3. Spatial light modulators and holography: how to play the notes back one at a time
4. How holographic optogenetics is manufacturing feeling
Live imaging of calcium indicators reads single-neuron activity in the brains of awake and behaving animals
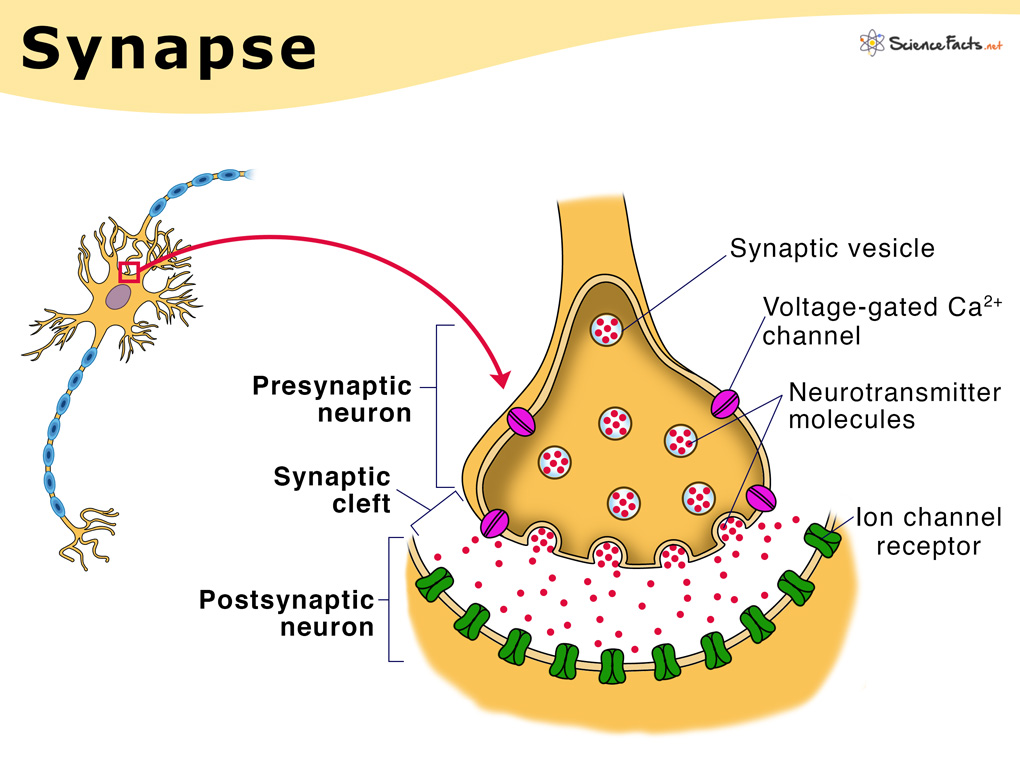
To grasp the principle behind calcium imaging, you must first know the basics of neurotransmission. In the brain, neurons communicate at sites called synapses (figure 1). These sites are located on a part of the neuron called the dendrite. They consist of a presynaptic terminal which is the conveyor of information, and the post-synaptic terminal, which is the recipient of information. The substrate of information used in synaptic transmission are molecules called neurotransmitters. At the postsynaptic terminal, receptors for these molecules transduce the chemical release into a change in the neuron’s electrical potential (figure 1). At the presynaptic terminal, neurotransmitters are packed into tiny vesicles which are exocytosed onto the postsynaptic neuron. Vesicle release is triggered by the release and concentration of calcium which acts as a second messenger to mobilize and exocytose vesicles (figure 1). This transient change in calcium concentration can be measured and used as a proxy for neural transmission.
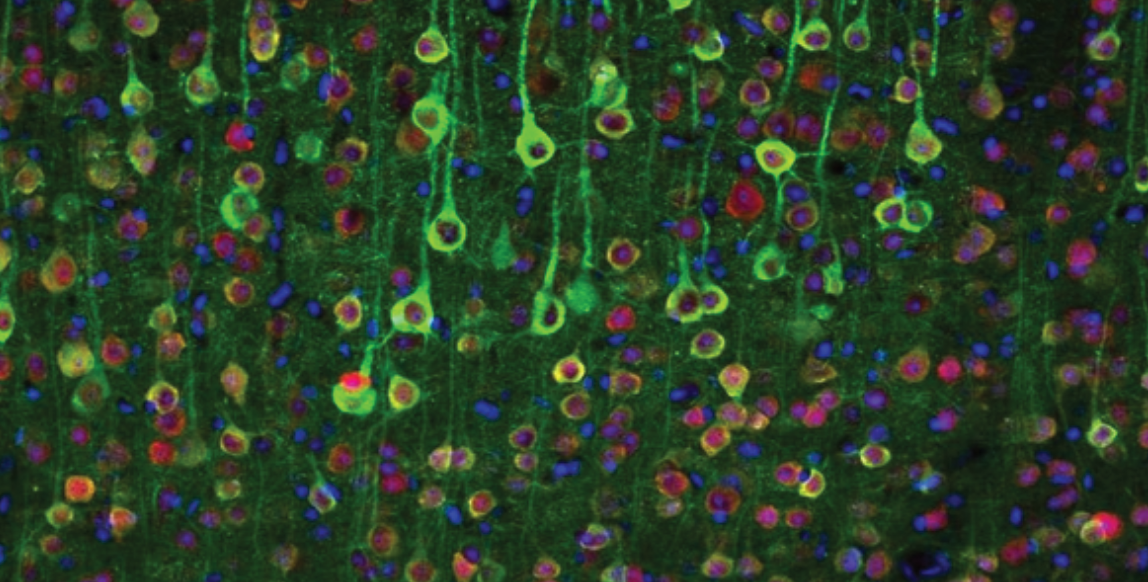
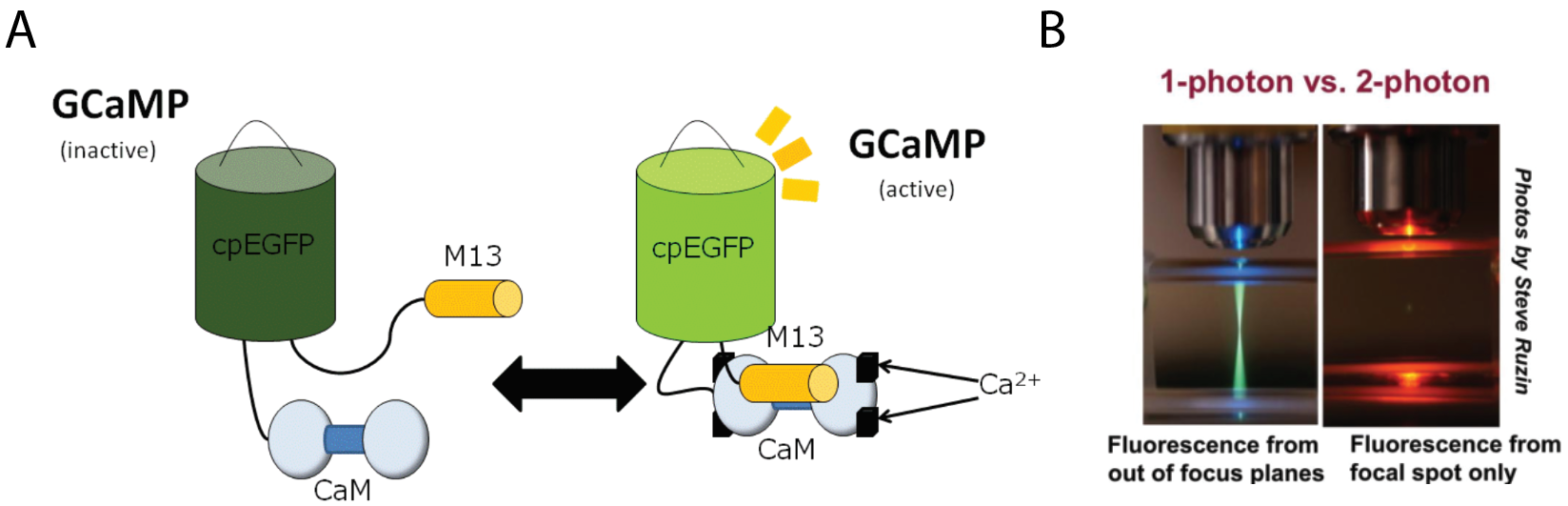
Calcium indicators capitalize on these cellular processes to report the activity of the neuron by optically outputting the change in calcium concentration in the cell over time. Calcium binds these indicators which causes a conformational change that alters the fluorescent properties of the protein (figure 2a). Using transgenics, scientists can express GCaMP (the most widely used calcium indicator) in genetically specified neural populations. The activity of these neurons can then be imaged with conventional optical techniques. More sophisticated imaging methods such as two-photon microscopy improve the spatial resolution of imaging which allows for improved imaging resolution and depth (figure 2b). These preparations can be used to image the activity of hundreds of neurons in live animals.
Optogenetics allows for reliable and temporally precise activation of neurons using light
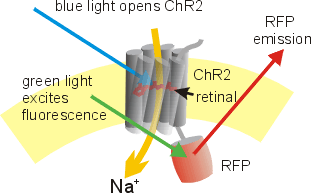
Now that we established a means of reading from neural circuits using calcium indicators, the second piece of the puzzle is to write back into these circuits. This is achieved using optogenetics. Optogenetics is a tool whereby a light-sensitive protein called channelrhodopsin is expressed in neurons. Channelrhodopsins can transduce a photonic signal into electrical activity. It does this by absorbing the energy from the light of an appropriate wavelength to induce a conformational change in the protein that enables it to conduct ions and shift the membrane potential of the neuron thereby causing neural firing (figure 3). Many channelrhodopsins have been discovered in nature and re-engineered for neuroscience. Compatibility with transgenics allows for localized expression in genetically identified neural populations. So, with a flip of a switch, you can turn specific neurons on and off. Optogenetics has revolutionized how neuroscience is conducted by offering a neat outlet to test the functional sufficiency of certain neurons.
Spatial light modulators effectuate an experimenter derived and spatiotemporally controlled holographic illumination pattern
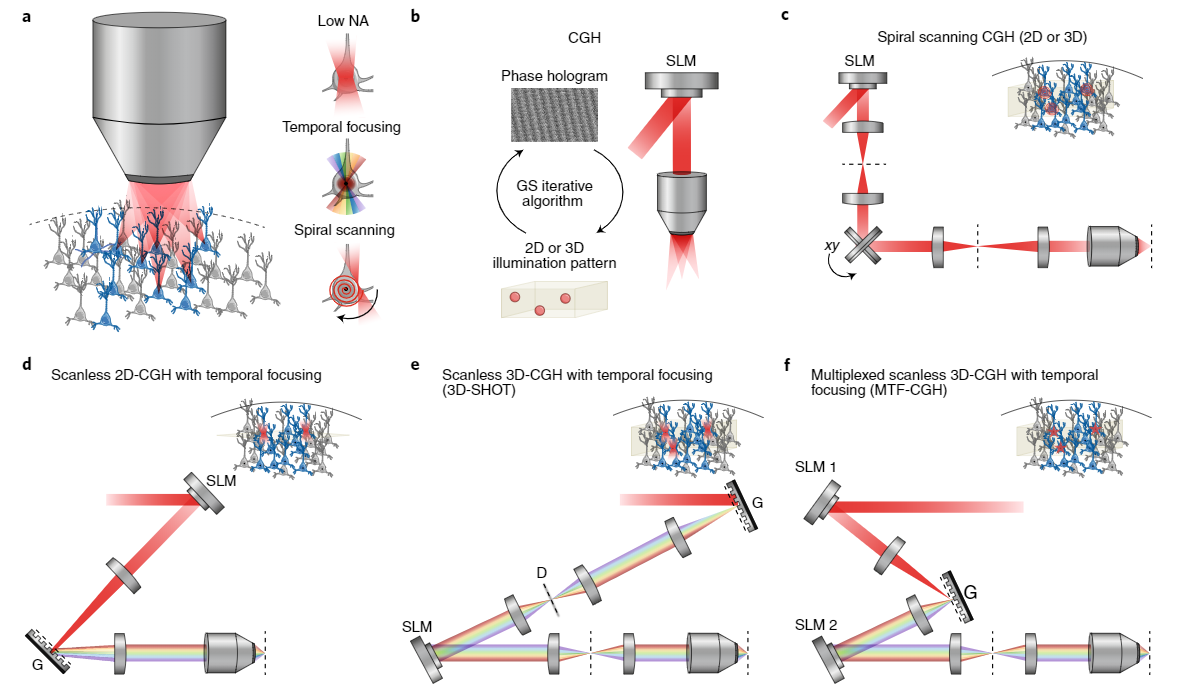
Writing into neural circuits with high spatial and temporal precision necessitates the generation of fine spatially localized holograms which can be programmed by reading the brain’s sheet music (see GCaMP imaging). These are generated using Spatial light modulators (SLMs) which use liquid crystal to modulate the phase and or amplitude of light. To achieve the 3D hologram, first, the phase hologram represents the optical interference pattern generated by the light emitted from an object and a perpendicular reference beam. The phase hologram can also be calculated using a Fourier transform-based iterative algorithm. The phase hologram can then be loaded onto the SLM to produce the three-dimensional illumination pattern (Figure 4a-b). Holographic optogenetic setups have been engineered using a multiplicity of approaches such as Spiral Scanning, Scanless 3D-CGH with temporal focusing (3D-SHOT), and Multiplexed scanless 3D-CGH with temporal focusing (MTF-CGH) (figure 4c-f). Temporal focusing allows high axial precision which enables activation of units within an individual plane.
For read/write capability, these custom-built microscopes will possess an imaging laser to record neural activity and a photostimulation laser to activate these circuits. How, though, do we equip cells with the right indicators and opsins such that we can have independent insight and access into their activity?
A Bicistronic virus infects cells with the genetic information for disparately activatable indicator and opsin
To perform the reading and writing operations separately, these lasers must emit light at differing wavelengths. Additionally, this necessitates calcium indicators and channelrhodopsins that are activated at these differing wavelengths. In recent work, scientists from Karl Deisseroth’s lab used GCaMP6m as a calcium indicator and ChRmine which is a potent channelrhodopsin. These can be activated using different wavelengths of light. The genetic information for both proteins was engineered into a bicistronic vector. This allows for the expression of both proteins from a single mRNA transcript. This vector can then be packaged into a virus and injected into a specified region of the brain where it can then infect cells to deliver the vector. These cells will now be equipped to translate the mRNA transcript into both the calcium indicator and channelrhodopsin proteins. How are scientists using this technology to advance our understanding of brain circuits?
Directly testing how ensembles modulate brainwide activity and mediate perceptually driven behavior using holographic optogenetics
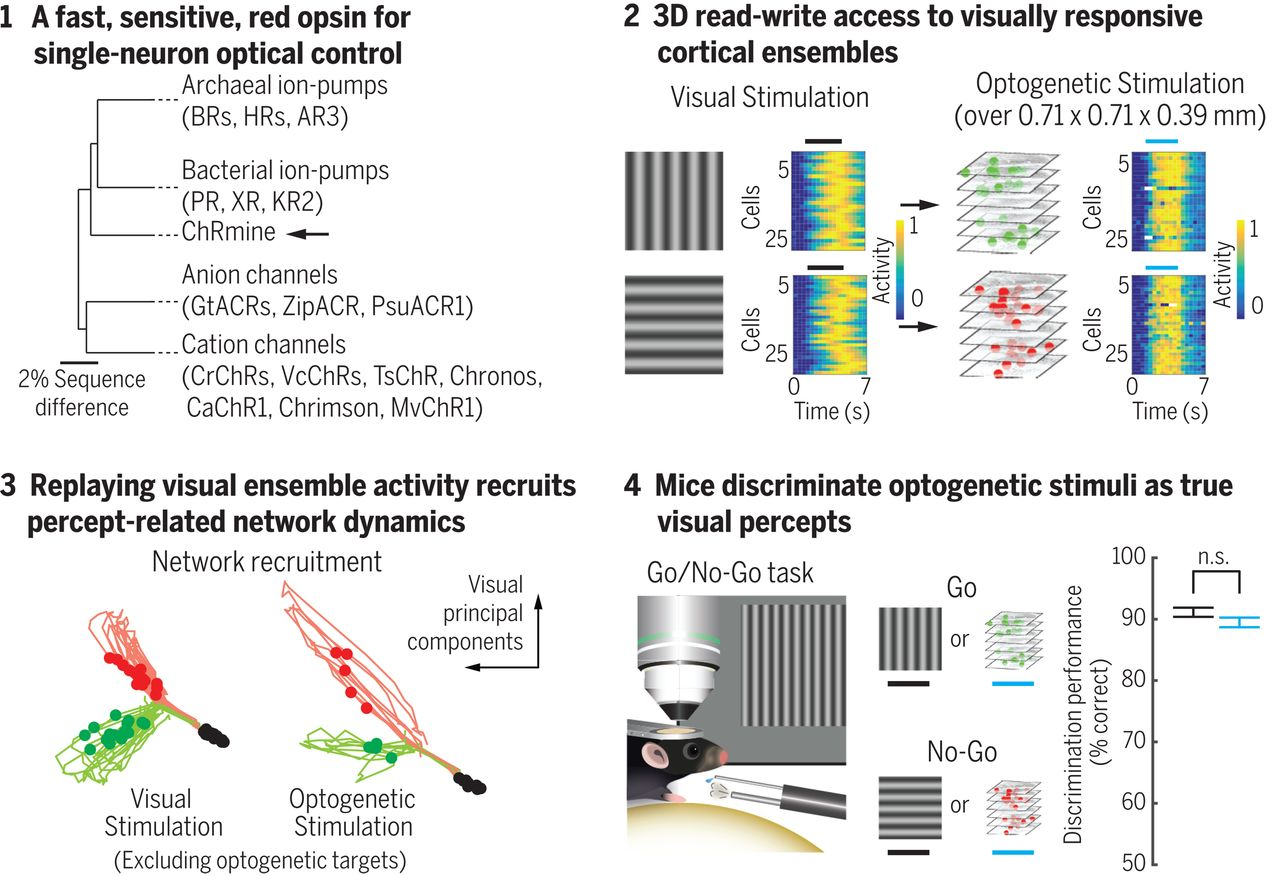
Holographic optogenetics, although a novel technology, is already proving useful. Preliminary work in the field demonstrated the potential for holographic optogenetics-mediated restoration of vision loss. The principal strategy behind their work was to activate precise populations of retinal ganglion cells in blind retinas to simulate natural patterns of visual activity. They were successful in generating rhythmic patterns of activity which mirrored activity dynamics when watching a visual scene of a running cat. Although this study was able to simulate patterns of activity, it was still unclear how this influenced total network dynamics and or if this activity could engender a perceptual change. More recent work from Karl Deisseroth’s lab approached these questions.
Scientists were able to read from neurons in the visual cortex to determine which cells would respond to the perception of vertical vs. horizontal visual gratings. After identifying vertical-preferring and horizontal-preferring neurons, they were able to selectively access these populations optogenetically. The artificial activation of these circuits recapitulated similar network dynamics to literally perceiving the visual stimulus. Remarkably, activating these individual populations caused the mouse to behave in a way that reports that it perceived a grating that was not there. In other words, writing into the appropriate circuits seemed to plant an artificial perception that was acted upon.
Obtaining the calcium transients of individual neurons includes acquiring the ROI of somas in imaging. Many algorithms have been developed to semi-automate this process and to extract single-neuron activity such as CNMF-E. This typically requires the identification of neurons across 2D image stacks. If you are interested in the automated identification of fluorescent ROIs such as somas, Biodock can help design and implement a custom AI module for this purpose! For further information, contact us to examine all the services Biodock has to offer, and for a consultation.
Conclusions: outlooks on how far we have come as inspiration for where we might go
Holographic optogenetics uniquely ties the worlds of microscopy, engineering, genetics, molecular biology, and systems neuroscience. It is inspiring to think that just forty years ago David Hubel and Torsten Wiesel received the Nobel prize for their pivotal work in identifying the feature detection properties of visual neurons. One at a time, they listened in on the electrical activity of individual neurons of the striate cortex in anesthetized cats as they presented them with a variety of visual stimuli. Four decades later, scientists can identify this same activity in hundreds of cells and play it back to recreate a perception possibly similar to that all those years ago. Who knows where the next four decades will take us.